6.3. Select menu
It is a menu related to the function of selecting atoms or molecules.
Hint
How to select atoms
There are two ways to select an atom: The method using a red circle marker and the method using a blue circle group selection.
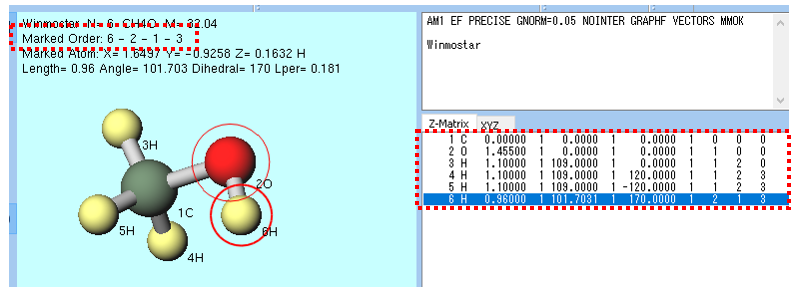
The method using the red circle marker is mainly used for operation on one atom.The marker moves to the atom that was left-clicked in Viewport.In the example of the methanol molecule in the figure below, the oxygen (red) and hydrogen (yellow) of the OH (hydroxy) group are left clicked. The red thick circle is drawn to the atom to which the marker is finally attached, and the red thin circle is drawn to the atom attached the marker one before.The marker also moves by left clicking at Coordinate Viewer in the right red frame of the figure below.Last four atoms marked are internally stored and displayed as shown in the upper left red frame of the figure below.The number of each atom can be confirmed in Viewport by selecting or Coordinate Viewer.
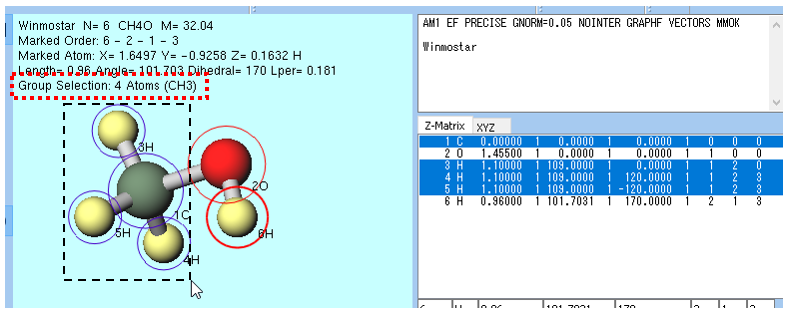
The method using group selection of the blue circle is mainly used for operations on multiple atoms.You can select a group by Ctrl+left drag , Ctrl+left click or Shift+left click in Viewport.You can also select a group by using the function of menu.In the example of the methanol molecule in the figure below, after Ctrl+left drag around the CH3 (methyl) group, it can be confirmed that the CH3 group atoms are surrounded by blue circles.You can also select a group by Ctrl+left click in Coordinate Viewer.On the upper left of Viewport, the number of atoms and composition of the group selected are displayed
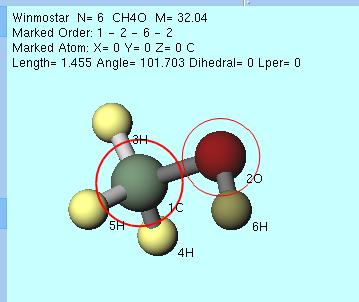
Select Partial Structure Split by 2 Marked Atoms is executed automatically when you execute an operation on multiple atoms such as in a state where no group is selected. Specifically, the substructures that are divided by two atoms with a red thick circle and a red thin marker attached are selected as a group. In the example of a methanol molecule shown below, after oxygen (red) and carbon (green) are moved in the order of left click and the marker is moved, Rotate group on axis (selection 2 atoms, mouse control) is selected, the group with the last marker on the CH3 group is automatically selected (highlighted on the screen).
6.3.1. Select All
Select all atoms as a group.
6.3.2. Select None
Cancel the group selection.
6.3.3. Invert Selection
Select a group that has not been selected as a group, and cancel the group selection of the group that was selected until then.
6.3.4. Select by Molecular Species
It opens with Use List tab of Select by window checked Molecular Species.Click on each line in the list in the window and the corresponding molecular species are selected.
Note
When calculating from protein pdb files, this function can be used to extract proteins, ligands, bound water, buffers, etc.
6.3.5. Select by Molecules
It opens with Use List tab of Select by window checked Molecules.Click on each line in the list in the window and the corresponding molecule is selected.
6.3.6. Select by Elements
It opens with Use List tab of Select by window checked Elements.Click on each line in the list in the window and the corresponding element is selected.
6.3.7. Enter Selection Language
It opens with Use Selection Language tab of Select by window.If you click on Apply button after describing in the text box, the corresponding atoms are selected.The following syntax is supported in the selection language.
Basic syntax
element C HSelect all carbon and hydrogen atoms.
index 1-3 6 8Select 1, 2, 3, 6 and 9th atoms.
compid 1
moleid 1-3
site 1
charge 0.5
resname GLYSelect the atoms whose residue name is GLY.
name CASelect the atoms whose atom name is CA.
x< 10Select the coordinates where the x-coordinate is less than 10 Angstrom. (No space between x and <.) Similarly, y and z can be used. You can also use
x>,x=>andx<=.
bondto HSelect the atom to be bonded to the H atom.
group mygroupSelect the registered group named “mygroup.”
The residue name and atom name can be used when PDB or gro format file is opened.
Composite syntax using logical operators
(compid 1) and (site 1)It is a molecular species whose CompID is 1 and the first site in the molecule is selected.
(current) and (element H)Select only hydrogen atom among atoms currently selected in the group.
(resname GLY) and (not (element H))Select the atom whose residue name is GLY and other than hydrogen.
In addition to
and,notyou can useorandxor. When using these logical operators, use the parentheses()as in the example above.Also, if you enter
%2==0where you enter an integer, it will assign a sequence of numbers where the remainder divided by two is zero. For example, typingsite %3==1will select atoms with sites 1, 4, 7, 10… will be selected.
6.3.8. Select Partial Structure Split by 2 Marked Atoms
Substructures that are separated between two atoms with the first and second markers are selected. Details are described in Select menu with pictures.
6.3.9. Group selection of atoms bound to the current group
Selects an additional group of atoms bound to the current selection group.
6.3.10. Group selection of atoms adjacent to the current group
Select a group of atoms adjacent to the current selection group. Here, adjacent atoms are defined as those atoms that have the same Voronoi boundary when Voronoi splitting is applied to all atoms.
6.3.11. Group selection of molecules adjacent to the current group
Select a group of molecules adjacent to the current selection group. Here, adjacent atoms are defined as atoms that have the same Voronoi boundary when Voronoi splitting is applied to all atoms.
6.3.12. Select Atoms Around Marked Atom
Select a group of atoms that exist within a specified distance from the marked atom.
6.3.13. Select Molecules Around Marked Atom
Selects a group of molecules whose center of gravity is within a specified distance from the marked atom.
6.3.14. Select Residue with Marked Atom
Select Residue with Marked Atom. Residues with the same residue number and consecutive atom IDs are determined to be the same residue.
6.3.15. Select Groups by Enlarging Each Molecular Species
Enable group selection of atoms by zooming in on the molecule in 3D for each molecular species. In the Create Group window, use Target to switch molecular species and Ctrl+click on the left side of the window to select atoms (sites). Click the Add button to register the selected sites, and click OK to create a group for each molecule’s registered sites. If Apply to First Molecule Only is checked, the operation is applied only to the first molecule of the selected molecular species.
6.3.16. Register Selected Group
Name and register the selected group so that it can be called from Select Registered Group.
6.3.17. Select Registered Group
Call the group registered with Register Selected Group. When LAMMPS is executed, the groups listed here are output in the ndx file. If you want to take over the registered group even after restarting Winmostar, export to the ndx file using Export Groups to Index File (ndx) and then read it using Import Groups from Index File (ndx).
6.3.17.1. Rename the registered group
Rename the group registered with Register Selected Group.
6.3.17.2. Delete the registered group
Delete the group registered with Register Selected Group.
6.3.18. Import Groups from Index File (ndx)
Reads the group output to the ndx file and makes it accessible from Select Registered Group.
6.3.19. Export Groups to Index File (ndx)
Outputs the groups registered in Select Registered Group to the ndx file.
6.3.20. Add Selected Group to Index File (ndx)
Add the group selected group to the specified ndx file.
6.3.21. Concatenate and Transpose Groups
This function combines multiple groups with equal number of atoms in them and transposes the order. For example, if group A contains 1, 2, 3, and group B contains 4, 5, and 6, and this function is applied to groups A and B, three groups will be created: 1 4, 2 5, and 3 6.